Glia Research Laboratory
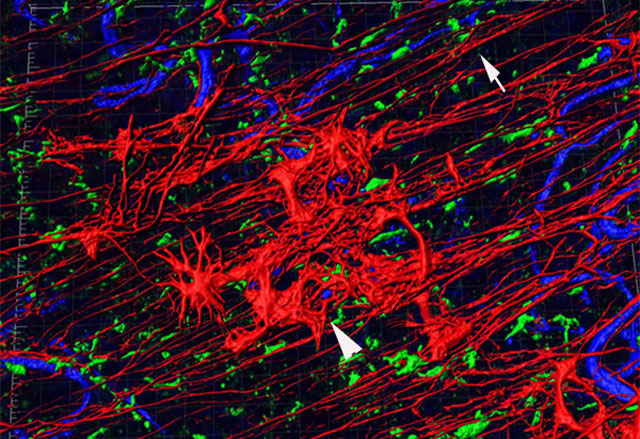 This 3D rendered image of a human retina shows glial cells in their normal position (arrow) and on the retinal surface, creating a bloom-like structure (arrowhead). Two different types of glia are shown, one labelled in red and the other in green as well as retinal blood vessels (blue). As shown, glia ensheath retinal vessels.
This 3D rendered image of a human retina shows glial cells in their normal position (arrow) and on the retinal surface, creating a bloom-like structure (arrowhead). Two different types of glia are shown, one labelled in red and the other in green as well as retinal blood vessels (blue). As shown, glia ensheath retinal vessels.There are two primary glial cells in the retina, astrocytes and Müller cells. These cells are not only crucial for maintaining normal retinal function, but are among the first to respond during injury or disease. Müller cells span the entire retinal thickness and make contact with all retinal cells. These glial cells become activated and can change structure, or remodel, in retinal diseases. Despite their early response to injury, the role of glial cells has only recently gained attention in the fight to understand retinal disease.
The Edwards lab, led by Malia Edwards, Ph.D., has recently identified glial changes in age-related macular degeneration (AMD). We have observed glial cells abnormally migrating onto the vitreo-retinal surface and into the subretinal space in eyes with advanced AMD. Glial migration onto the vitreo-retinal surface is most prominent in eyes with neovascular AMD while subretinal migration is more striking in eyes with geographic atrophy. The Müller cells are also activated and disorganized throughout the retina in AMD, suggesting that their normal function may be altered. Our research is focused on understanding the glial changes seen in AMD, with three main focal areas:
- How early in the disease process are Müller cells affected?
- What stimulates Müller cell remodeling and membrane formation in AMD?
- How glial remodeling, activation and membrane formation affect AMD disease progression and treatment efficacy?
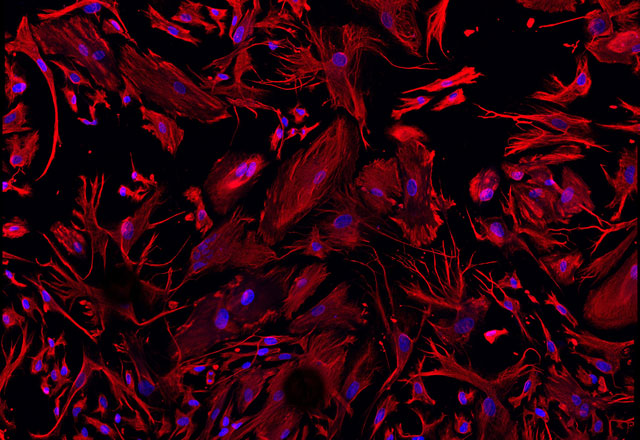 Our research also involves using primary Müller cells. Shown here are Müller cells, isolated from a human retina, stained with vimentin (red) and DAPI (blue). These primary cells allow us to directly determine how different factors present in AMD influence Müller cells directly
Our research also involves using primary Müller cells. Shown here are Müller cells, isolated from a human retina, stained with vimentin (red) and DAPI (blue). These primary cells allow us to directly determine how different factors present in AMD influence Müller cells directlyWhile the primary focus of my research has been AMD, we have also observed extensive Müller cell remodeling and membrane formation in other retinal diseases, including Stargardt disease and retinitis pigmentosa. Therefore, understanding the causes and consequences of Müller cell remodeling in AMD may help us understand and treat other retinal diseases as well.
The Edwards lab is interested in understanding the roles Müller cells and astrocytes play in both pathological and normal vascular development. During my post-doctoral training, I discovered the importance of Lama1 in retinal vascular development. Interestingly, mice with mutations in Lama1 develop membranes similar to those seen in aged human eyes. These mice may serve as a model for understanding glial membrane formation and/or studying how we can reduce membrane formation.
Principal Investigator
Contact Info
M023 Smith Building
410-614-9888
email: [email protected]
Recent Publications:
- Bhutto IA, Ogura S, Baldeosingh R, McLeod DS, Lutty GA, Edwards MM*. An Acute Injury Model for the Phenotypic Characteristics of Geographic Atrophy. IOVS 2018; Mar 20; 59(4):AMD143-AMD151.
- Goldberg M, McLeod DS, Tso M, MD, Packo K, Edwards M, Bhutto IA, Baldeosingh R, Eberhart C, Weber B, Lutty GA. Ocular Histopathology and Immunohistochemical Analysis in the Oldest Known Individual with Autosomal Dominant Vitreoretinochoroidopathy. Ophthalmology Retina 2017; 1-19.
- Edwards M*, McLeod DS, Bhutto I, Grebe R, Duffy M, Lutty G. Subretinal Glial Membranes in Eyes With Geographic Atrophy. IOVS 2017 Mar 1;58(3):1352-1367.
- Edwards MM*, McLeod DS, Bhutto IA, Villalonga MB, Seddon JM, Lutty GA. Idiopathic preretinal glia in aging and age-related macular degeneration. Exp Eye Res. 2016; Sep;150:44-61. PMCID: PMC4728061
- Edwards MM*, Rodríguez JJ, Gutierrez-Lanza R, Yates J, Verkhratsky A, Lutty GA. Retinal macroglia changes in a triple transgenic mouse model of Alzheimer's disease.” Exp Eye Res. 2014; Oct; 127:252-60. PMCID: PMC4175519
- Edwards MM, McLeod DS, Li R, Grebe R, Bhutto I, Mu X, Lutty GA. The deletion of Math5 disrupts retinal blood vessel and glial development in mice. Exp Eye Res. 2012; Mar;96(1):147-56. PMID: 22200487. PMCID: PMC3296879
- Edwards MM*, McLeod DS, Grebe R, Heng C, Lefebvre O, Lutty GA. Lama1 mutations lead to vitreoretinal blood vessel formation, persistence of fetal vasculature, and epiretinal membrane formation in mice. BMC Dev Biol. 2011; Oct 14;11:60. PMID: 21999428. PMCID: PMC3215647
- Edwards MM, Lefebvre O, Jordan WL, Smith RS, Wu J, Peachy NS, Naggert J, and Nishina PM. A mutation in lama1 disrupts retinal blood vessel development and inner limiting membrane formation. JBC. 2010; March; 285(10): 7697-711. PMCID: PMC2844215PMCID: PMC2891478.

