Knowledge is the key to understanding restless legs syndrome (RLS). At the Center for Restless Legs Syndrome, established by Drs. Christopher J. Earley and Richard Allen, we conduct research to learn more about this disorder and deliver better care to people who live with it. Our care program focuses on patient education and includes standard-of-care treatments and research protocols not available in other centers.
Our Restless Legs Syndrome Experts
Hear From Our Experts
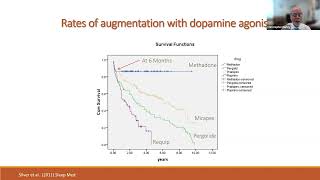
Biological Basis for and Management of Drug Induced Symptom Augmentation in Restless Leg Syndrome
Altered Iron Regulation: A Potential Cause of Restless Leg Syndrome
Iron and Restless Legs Syndrome
Role of Opioids in Restless Leg Syndrome
RLS Clinical Trials
Every day, research uncovers new information about possible therapies for RLS. Your involvement in clinical studies could help in the development of medications to treat RLS. You, a family member, and many other people may benefit from your willingness to become involved.
Learn about importance of clinic research
Understanding the Role of Epigenetics in RLS
This study is designed to address the question of why RLS has such a high inheritance risk. To date, there has been no single gene identified that can account for the 35-40% relative risk of inherency. This study looks deeper into complex genetic regulation (epigenetics) to determine if inheritance risk lies within these factors.
- Eligibility: women who have iron deficiency anemia. We are looking for women who do and do not (control group) have RLS symptoms.
- Contact: 410-550-1046
RLS Research Areas
Research is a very important component of our center. Knowledge is the key to better understanding this disorder and coming to terms with what we do not know about it. It is through research that we will hopefully one day bring about a cure for RLS.
-
Gaining a more thorough understanding of the genetic underpinnings of RLS should eventually help us to better understand the underlying pathophysiology of RLS. This should therefore also help us to develop more sophisticated and personalized methods of treatment.
Our research group was already involved in a Genome-Wide Association Study, or GWAS. This method screens for natural, single-unit variances in the DNA structure (like looking for a small difference in a single pearl compared to all the other pearls that make up a necklace). To date, several genes using the GWAS technique have been identified. These genes have one or more small alterations in their DNA structure that are associated with increased risk of developing RLS. These small changes are not “causing” the disease in the same sense that mutations in clotting factor genes cause bleeding problems in a hemophilia. These small differences in the genes have, in turn, small subtle effects that when added across several genes may then create a greater risk. This means we need to understand the dynamic relationship between these different genes. Even though these genes now appear to have little or no relationship to each other, they all relate to RLS. We are working on discovering what those dynamic relationships are by better understanding the biological roles that each of these genes play within cells. If we can establish a relationship between these RLS-related genes and their protein products, then we can begin to understand the specific biological mechanisms that are unique to RLS. From that we can develop potential interventions that might be used to change that system in a positive and personalized way.
In a large family history study, we demonstrated that the risk of inheritance looked very much like it was related to a dominant gene. But despite much effort and multiple studies, a single dominant gene has not been found. The genes found using GWAS only provide about 7% of the overall genetic risk and do not account for the dominant nature of the inheritance of this disease. Our group at Johns Hopkins has embarked on a new approach to genetic analysis, which involves looking, not just at the gene, but also at the multi-level factors that control how a gene is eventually transcribed into a protein. Certain environmental triggers can change how a gene is activated by activating complex biological processes referred to collectively as “epigenetic” mechanisms. These epigenetic mechanisms place (or remove) the equivalent of a small 'tag' on the gene(s) that may then alter the gene’s expression. These epigenetic changes can pass from one generation to another. Thus an epigenetic approach may provide the elusive answer as to what is inherited across family members with RLS.
-
Conditions in our environment can irreversibly change how and what genes are activated, a process referred to as epigenetics. It makes intuitive sense that an individual with a fixed set of genes should be able to modify expression of those genes based upon the individual’s environmental challenges. Unlike genetic changes, which are fixed, epigenetic changes usually occur under environmental stress and thus could be prevented or even reversed if we can identify the stressors or the genes that have been modified. The results from our large family study identify “environmental factors” as contributing more to the risk of developing RLS than that from just genes alone. The question is what are the environmental factors? We have identified iron insufficiency as the leading environmental factor contributing to the development of RLS. Whether iron insufficiency develops during childhood, pregnancy, or throughout the lifetime of an individual, its consequences may be permanent even if the iron levels are restored to normal. Our research has led to the discovery of altered brain iron regulation as a key component underlying RLS. We believe the alteration in brain iron regulation is a consequence of this environment-gene interaction. We are now striving to understand epigenetic and genetic factors regulated by environment stressors such as iron insufficiency and how they can permanently change brain iron regulatory mechanisms. It is important to identify the nature of the environmental stressor (duration, severity and timing) and the genes that are affected in order to potentially prevent or change the outcome, which is the development of RLS symptoms.
-
We use animal and cell models that represent aspects of what we currently know about genetic, epigenetic and other biology mechanisms associated with RLS. By utilizing these models, we can gain an understanding of how certain genes relate to other genes and the consequence of this dynamic gene interaction, particularly as it relates to brain regulation. We can explore the consequence of environmental stressors like iron deficiency at different ages of the animals and then look for epigenetic changes within the RLS-related genes or in genes not yet identified. In the models, we will be able to pose questions, collect data, and grow a tangible, viable, and operationally significant dynamic system of the underlying biology and potential pathophysiology behind RLS. Our models provide, for the first time, a chance to actually test new compounds for this disease.
Support RLS Research and Care at Hopkins
Philanthropy is essential to our continued progress toward improving the lives of people with RLS. The generosity of individuals enables our work to continue even when federal grant funding falls short. Gifts can be designated to support specific research projects or left unrestricted, to support either a particular physician’s work or RLS research in general. These gifts help us address new opportunities and unexpected needs quickly.
We appreciate your support and consideration. To learn more about Dr. Christopher Earley or our center and its activities, please contact:
Angel Terol
Senior Associate Director of Development
Fund for Johns Hopkins Medicine
Department of Neurology and Brain Sciences
[email protected]
(443) 287-7873