Study Shows HIV Remission Is Possible for Children Started on Very Early Antiretroviral Therapy
Read Full ArticleLatest News
-
04/16/2024
Study Suggests Adolescent Stress May Raise Risk of Postpartum Depression in Adults

-
04/12/2024
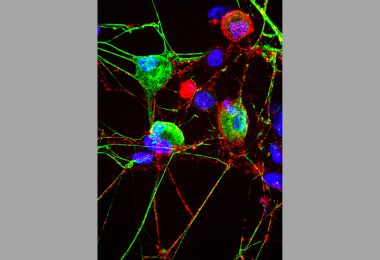
Cell’s ‘Garbage Disposal’ May Have Another Role: Helping Neurons Near Skin Sense the Environment
-
04/10/2024
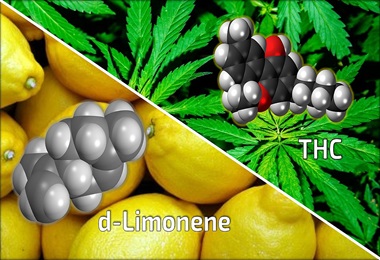
Researchers Show Chemical Found Naturally in Cannabis May Reduce Anxiety-Inducing Effects of THC
Contact Johns Hopkins Media Relations
Johns Hopkins Medicine In the News
-
What to do on the nights you are struggling with insomnia, according to experts
If you haven’t fallen asleep in roughly 15 to 20 minutes, go into another room and try another activity until you start to feel drowsy and try again, says Dr. Rachel Salas, professor of neurology at the Johns Hopkins University School of Medicine.
-
Bridging the gap between medicine and community health
Through the Community Health Workers program at Johns Hopkins Howard County Medical Center, about 100 people have trained to become community health workers. It's 13 weeks of training where they become a jack of all trades, learning everything from CPR to information on health screenings.
